GTax¶
Introduction¶
GTax python package provides tools for the creation of the GTax sequence-based database. This database includes one assembly per organism deposited in the NCBI Genomes database. The sequences are organized by 19 taxonomic levels from superkindom to clades. Python pickle files are also provided with a graph data structure for the taxonomic tree.
GTax is comprised of 19 taxonomic levels that cover all taxonomic superkingdoms:
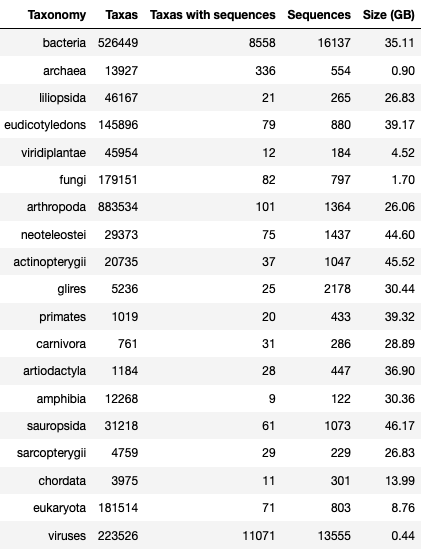
Current version¶
The current public version is available in GCP for download. You need to use your GCP project as this bucket uses Requester Pays option.
gsutil -u <you-GCP-project> -m cp -r gs://gtax-database/<latest_version> .
The database is comprised of two folder: blastdb and fasta. The blastdb folder include the BLAST indexes for BLAST searches. The fasta folder includes the FASTA files for the taxonomy groups and two Python Objects in pickle files (taxonomy.pickle and taxonomy_groups.pickle) which are used to load all GTax metadata into the [Taxonomy class](https://github.com/ncbi/gtax/blob/main/src/gtax/taxonomy.py#L37)
Help and Support¶
For query/questions regarding GTax, please write write veraalva@ncbi.nlm.nih.gov
For feature requests or bug reports, please open an issue on our GitHub Repository.
Public Domain notice¶
National Center for Biotechnology Information.
This software is a “United States Government Work” under the terms of the United States Copyright Act. It was written as part of the authors’ official duties as United States Government employees and thus cannot be copyrighted. This software is freely available to the public for use. The National Library of Medicine and the U.S. Government have not placed any restriction on its use or reproduction.
Although all reasonable efforts have been taken to ensure the accuracy and reliability of the software and data, the NLM and the U.S. Government do not and cannot warrant the performance or results that may be obtained by using this software or data. The NLM and the U.S. Government disclaim all warranties, express or implied, including warranties of performance, merchantability or fitness for any particular purpose.
Please cite NCBI in any work or product based on this material.